
Holocentrism
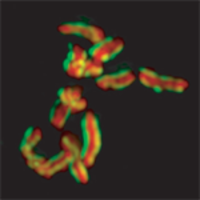
C. elegans (Holocentric) Oegema et al. Wormbook.org Cell division chapter
The Holocentrism project is a URGI genome dynamics research project whose goal is to investigate the molecular nature of holocentric centromeres by comparative and functional genomics in Insects, more particularly focusing on synteny, chromatin expression and genetic recombination.
URGI team: Hadi Quesneville, Emmanuelle Permal with the help of URGI support team
Introduction
Under the word holocentric, cytologists have described chromosomes which are devoid of any major centromeric constriction, so that the mitotic pairing occurs along almost the whole length of the chromosomes, and chromatids separate at anaphase as parallel bodies.
Holocentric biological diversity
Organisms with holocentric chromosomes can be found in both plant and animal kingdoms and may be the result of convergent evolution. Alternatively, the ancestral eukaryotic chromosome may have been holocentric and restriction of the centromere to a localized region has been a repetitive evolutionary event leading to monocentrics. Apart from Lepidoptera, holocentric chromosomes have been found in aphids, in nematodes like C. elegans, and in some wild plants (Juncaceae and Cyperaceae, the closest relatives of Graminaea).
Holocentric chromosome structure
Holocentric chromosomes differ from the monocentric ones in centromere structure. In C. elegans, during mitosis, kinetochore spans most of the chromosome length .
At meiosis, conventional electronic microscopy failed to show a trilaminar kinetochore structure. However confocal microscopy allowed visualization of kinetochore proteins (KNL-1, HIM-10) forming a cup-like structure that encloses the ends of the chromosomes. Although reverse genetics via RNA interference in C. elegans led to the discovery of more than 30 kinetochore proteins among which, HCP-3 and HCP-1, homolog to the human centromeric chromatin binding proteins CENP-A and CENP-C, DNA sequences interacting with them have not been identified yet. The observed staining pattern of anti-HCP3 (CENP-A) antibodies as puncta within interphase nuclei indicates that not all genomic sequences are involved in centromere.
From the whole genome sequence data, the distribution of the most abundant repeats - the miniature inverted-repeat transposable element (MITE) like repeats - shows i) a conserved distribution for each elements in the 5 autosomes, ii) a tendency for 3 out of 4 MITE-like element studied to cluster at the chromosomal arms. The distribution of many other repeats belonging to C. elegans genome has been shown to mostly display a random distribution except for one 24 bp repeat, called CeRep25B, which forms two arrays on chromosome III specifically. None of these repeats have yet been shown to be involved in centromeres.
Up to now, only very parcellar data are available on other species . The first lepidopteran satellite sequence described is a 234 bp sequence in the cabbage moth, Mamestra brassicae that makes foci in in situ hybridization on the sex chromosomes Z and W. In plants, the use of the rice 155 bp RCS2 centromeric repeat as probe allowed the isolation of a 178 bp tandem repeat from Luzula nivea (belonging to the Juncacae family) that is organized in at least 5 large clusters of heterochromatin distributed along each of the twelve chromosomes of Luzula. A centromere-specific histone H3 (LnCENH3) has also been identified in Luzula nivea that allowed visualization via immunostaining of the diffuse centromere-like structure along the chromosomes.
Thus, despite the abundant genomic resources available for C. elegans, very much remains to be done to define the structure and functioning of holocentric chromosomes . We believe that comparative genomics could help in filling the gap between structure and function: Insects – which are often holocentric species –are very good candidates for performing such analyses.
Cytologic observations in insects
The chromosomes of several insect orders separated by at least 300 millions years have been described as holocentric. Some chromosomal fragments obtained after irradiation (inducing chromosome breakage) can be maintained over several generations. Several authors have described holocentric chromosomes in meiotic cells (on testis or ovarioles of Vth instar insect larvae. Electronic microscope slides were published, which confirm the existence of kinetochore plaques extending over 50 % to 70 % of the chromosome pole ward surface of the chromosome. In a holocentric chromosome one expects that the key sequences elements would be a kind of mosaic among other coding or non-coding sequences, but up to now no one knows the molecular nature of holocentric centromeric sequences and their regulation.
Goal
The project aims at determining the global genomic organization of putative centromeric structures in 4 holocentric species belonging to either Lepidoptera or Hemiptera, by combining both a comparative (bio-informatic) and functionnal (biological experiments) approach. First, repeated sequences and TE will be searched in two whole genomes (silkworm and pea aphid) and in partial genomic fragments (two Noctuidae species). These analyses will culminate by comparative genomics approaches between insect orders and between species of the same Order (Lepidopteran). Second, Lepidoptera centromeric regions– for which preliminary data are already available - will be purified on the basis of centromeric protein affinity and characterized. Third, the effect of a dispersed centromere on gene expression will be investigated in the frame of a high epigeneticism responsible for changes in reproductive mode in the pea aphid. Finally, through collaboration with a Japanese group, we will investigate in Lepidopteran the consequence of holocentrism in genetic stability and genome recombination.
Four topics are investigated:
- Repeated Sequences, transposable elements and comparative genomics of holocentrism
- Characterization of centromeric proteins in holocentric chromosomes
- Heterochromatin, epigenetics and holocentrism
- Genetic stability and holocentrism
Duration: 01/01/2008 to 31/12/2010
Coordinator: Philippe Fournier
Partners
| Philippe | Fournier | Unité de Biologie Intégrative et Virologie des Insectes, INRA - UMII, Montpellier, France |
| Hadi | Quesneville | Unité de Recherche Génomique Info - INRA 1164, Versailles, France |
| Denis | Tagu | BiO3P INRA – Agrocampus35653, UMR 1099, INRA Rennes |
| Kasuei | Mita | Insect Genome Research Unit, Tsukuba, Japan |
Results have been published in:
Extensive synteny conservation of holocentric chromosomes in Lepidoptera despite high rates of local genome rearrangements. d'Alençon E, Sezutsu H, Legeai F, Permal E, Bernard-Samain S, Gimenez S, Gagneur C, Cousserans F, Shimomura M, Brun-Barale A, Flutre T, Couloux A, East P, Gordon K, Mita K, Quesneville H, Fournier P, Feyereisen R. Proc Natl Acad Sci U S A. 2010 Apr 27;107(17):7680-5. Epub 2010 Apr 13. PMID:20388903
Genome sequence of the pea aphid Acyrthosiphon pisum. International Aphid Genomics Consortium. PLoS Biol. 2010 Feb 23;8(2):e1000313. PMID:20186266
Correlation of LNCR rasiRNAs expression with heterochromatin formation during development of the holocentric insect Spodoptera frugiperda. Stanojcic S, Gimenez S, Permal E, Cousserans F, Quesneville H, Fournier P, d'Alençon E. PLoS One. 2011;6(9):e24746. Epub 2011 Sep 30. PMID:21980354
Creation date: 27 Apr 2010
 eZ Publish
eZ PublishPublication supervisor: A-F. Adam-Blondon
Read Credits & General Terms of Use
Read How to cite









